Programmable RRAM Probabilistic Hardware for Fast Molecular Docking in Drug Discovery
Executive Summary
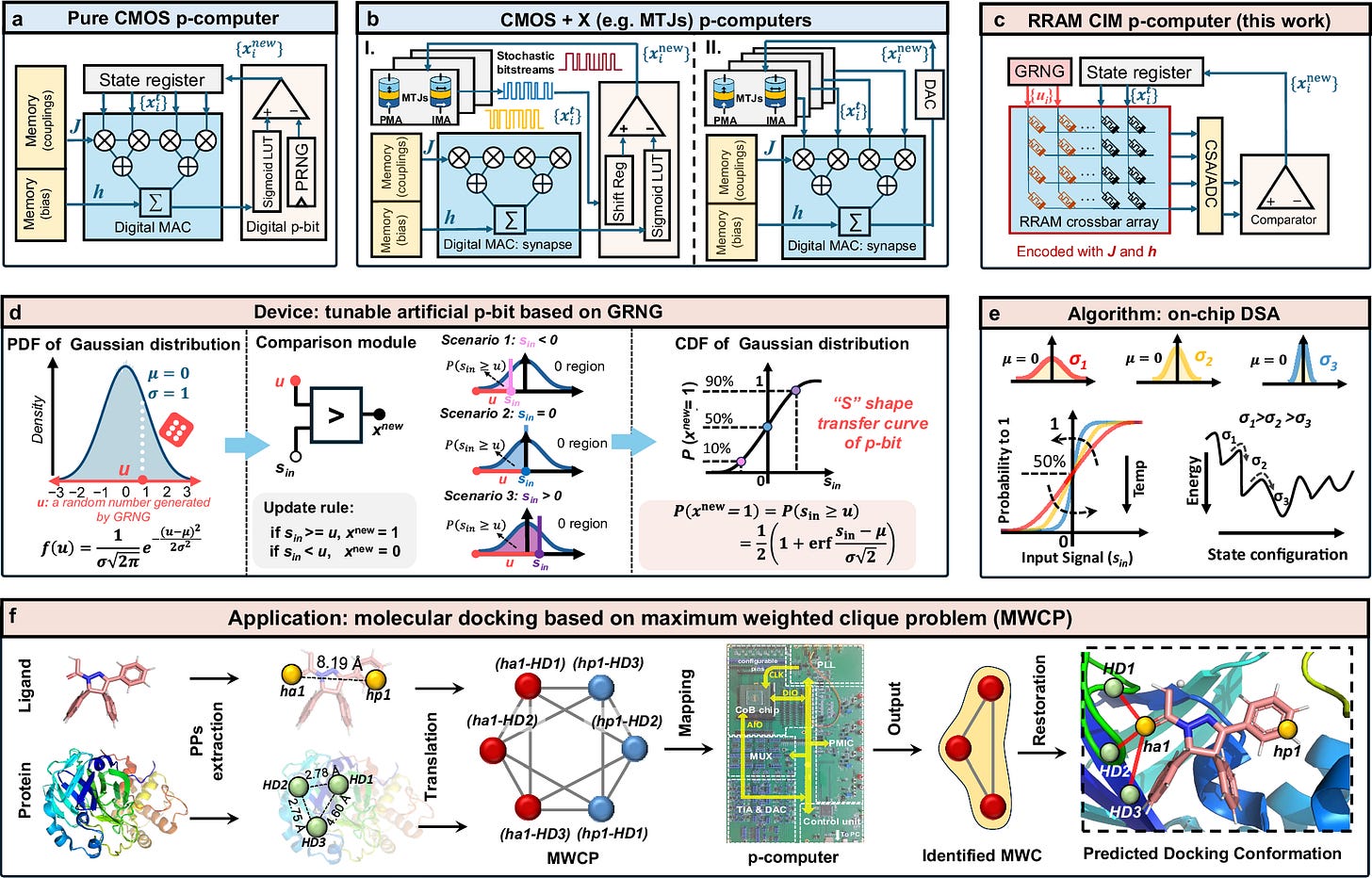
This technology demonstrates a hardware probabilistic computer designed to accelerate molecular docking in drug discovery. Molecular docking requires exploring vast combinatorial search spaces due to flexible ligand conformations and complex biomolecular structures. Conventional digital processors face memory bottlenecks and high energy costs when solving such combinatorial optimization problems. Quantum computing approaches show promise but remain constrained by scalability and hardware limitations. The presented system implements a probabilistic computer based on artificial probabilistic bits fabricated in 180 nm CMOS with back-end-of-line HfO₂ resistive random-access memory. The architecture integrates compute-in-memory functionality, directly encoding interaction weights within an RRAM crossbar array. Stochastic behavior is generated through a Gaussian random number generator that produces tunable sigmoidal probability responses. This design removes the need for external memory shuttling between processing and storage units. A dynamic slope annealing mechanism modulates stochasticity directly in hardware, enabling on-chip optimization. The system experimentally solved a 42-node molecular docking instance involving the LolA–LolCDE complex relevant to antibiotic development. Results were consistent with established protein–ligand interaction analysis tools. The prototype achieved high energy efficiency and reduced time-to-solution compared with conventional approaches. This work represents an early hardware realization of probabilistic computing for computational biology. It establishes a scalable semiconductor pathway for addressing complex bioinformatics optimization tasks.